ラボ
日本のラボ紹介
熊本大学ヒトレトロウイルス学共同研究センター 免疫・感染ダイナミックス分野
当分野では、免疫応答と感染応答を静的に分類するのではなく、時間の流れの中で理解することを重視しています。
とくに、T細胞における抗原受容体シグナル、Foxp3発現、免疫チェックポイント分子、サイトカイン応答の制御が、時間とともにどのように変化し統合されるかを、実験免疫学と計算解析の両面から研究しています。
蛍光Timerを用いたTocky系を基盤として、細胞の現在の状態だけでなく、直前までの刺激履歴や応答の推移を読み解くことを目指しています。

研究概要
免疫系は、感染や組織環境の変化に応じて、きわめて動的に応答します。
当分野では、このような免疫応答の時間的変化に注目し、細胞状態の遷移、分化、持続的刺激への適応、ならびに遺伝子発現制御の過程を明らかにすることを目指しています。
主な研究対象は、基礎免疫学、感染免疫学、胸腺におけるT細胞分化、Foxp3発現制御、がん免疫などです。
研究内容
T細胞受容体シグナルの時間動態
抗原刺激を受けたT細胞が、どの程度の強さで、どのくらい持続的に応答するかを解析します。
活性化の有無だけでなく、時間的に連続した応答過程として理解することを重視しています。
胸腺における分化と選択
発生途上のT細胞が、どの段階でどのような刺激を受け、異なる分化経路へ進むのかを調べます。
時間情報を導入することで、分化過程の理解を深めます。
Foxp3発現と免疫制御分子の動態
Foxp3発現、免疫チェックポイント分子、サイトカイン関連分子が、TCRシグナルと他のシグナル経路のもとでどのように誘導・維持・変化するかを解析します。
固定的な細胞分類ではなく、分子制御ネットワークの動態として免疫制御を理解することを目指しています。
感染・腫瘍環境における免疫応答
感染や腫瘍微小環境において、免疫細胞が時間とともにどのように変化するかを明らかにし、応答の成立と破綻を理解することを目指しています。
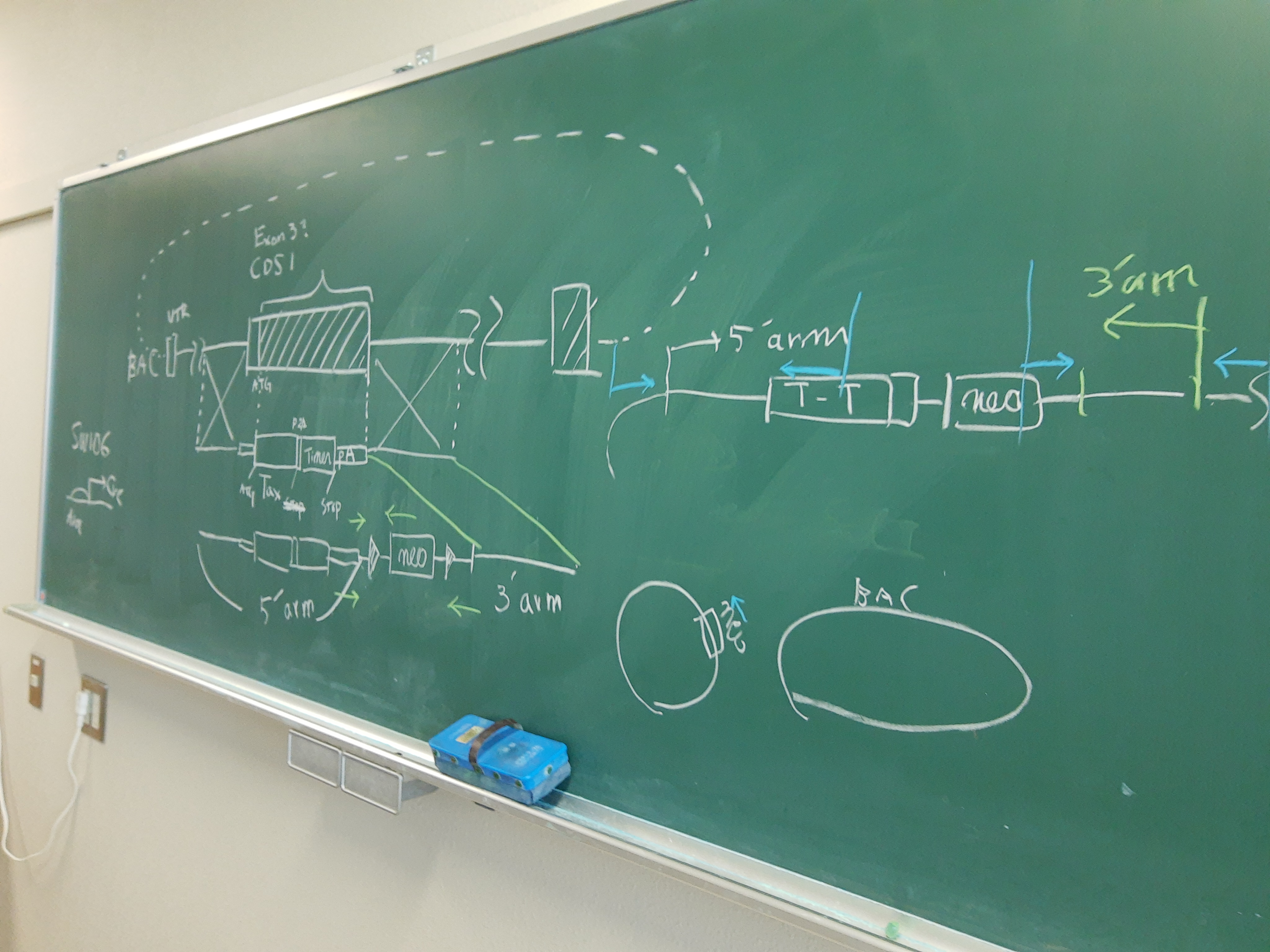
技術基盤
Tocky
蛍光Timerタンパク質を用いて、細胞内イベントの時間情報を可視化します。
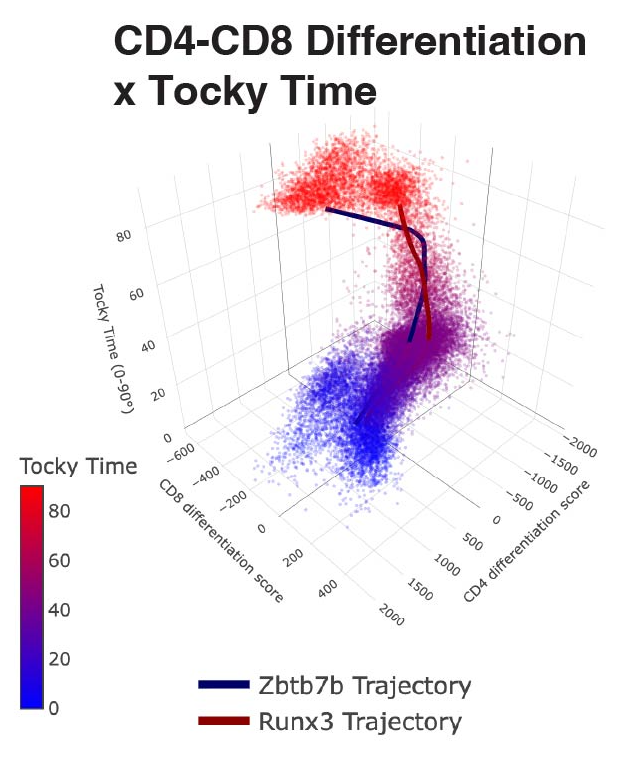
Figure from our bioRxiv preprint: Canonical Analysis of Fluorescent Timer-Anchored Transcriptomes Resolves Joint Temporal and Developmental Progression.
フローサイトメトリー
多色フローサイトメトリーを用いて、細胞集団の状態を高解像度で解析します。

単一細胞解析
単一細胞トランスクリプトーム解析・フローサイトメトリー解析などを通じて、細胞状態の遷移や多様性を解析します。
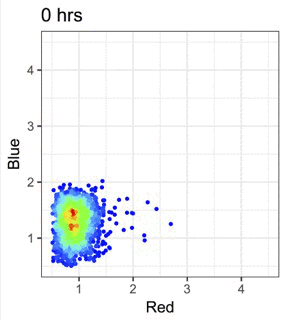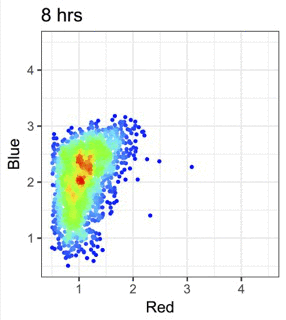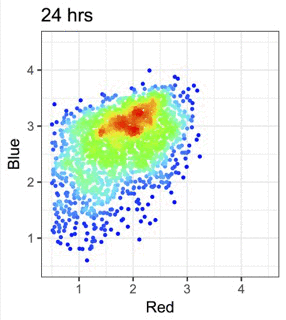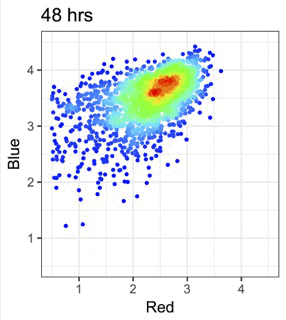
機械学習と免疫データ解析の融合
機械学習や多次元解析を活用して、Tockyデータやフローサイトメトリーデータの解析手法を開発しています。これらの手法を広く共有することで、研究基盤と解析環境の整備も進めています。
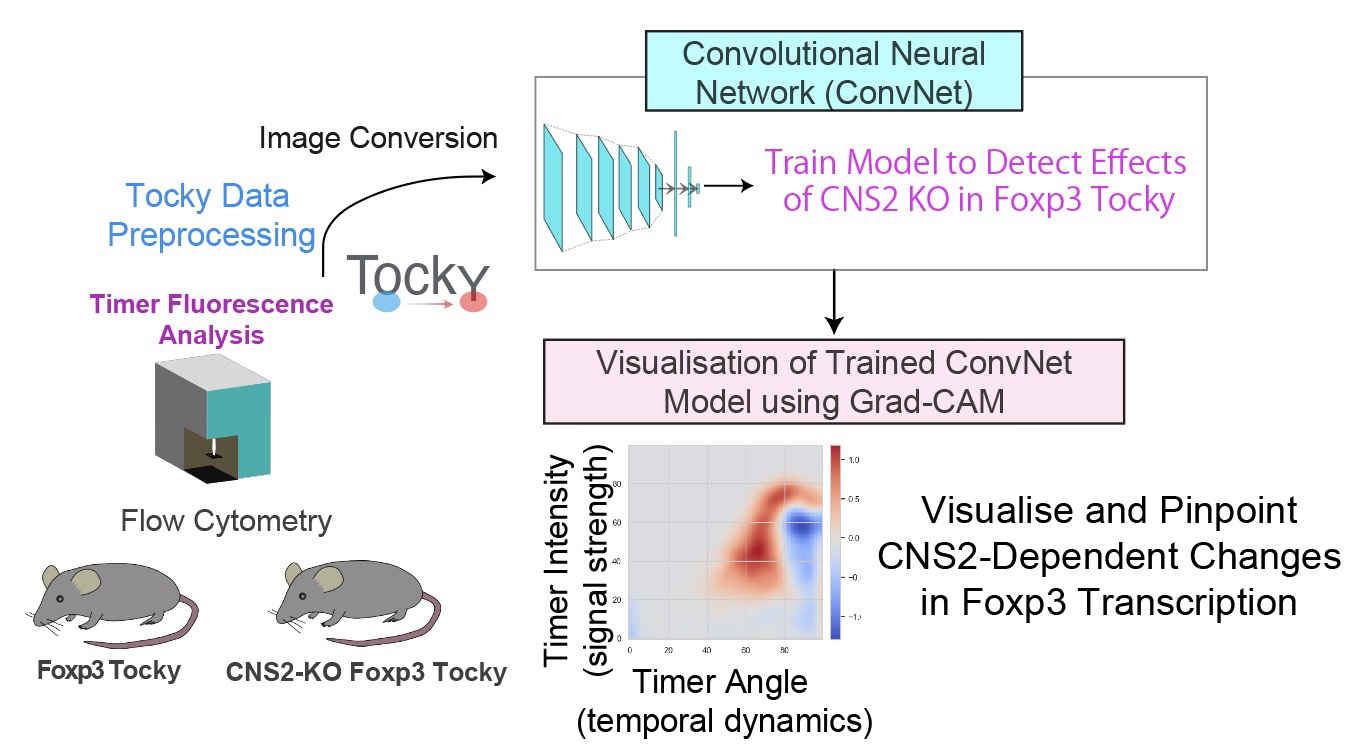
Figure modified from Irie et al., Nature Communications (2025): Machine learning-assisted decoding of temporal transcriptional dynamics via fluorescent timer.
共同研究
当分野では、免疫学、感染研究、フローサイトメトリー、単一細胞解析、計算解析を横断する共同研究を歓迎しています。
連絡先
news
| Mar 18, 2026 | 📄 New Preprint Published 🧭 Our new bioRxiv preprint introduces mCanonicalTockySeq, a framework that brings an experimental time anchor into single-cell transcriptomics to analyse temporal and developmental progression together. Using the Nr4a3-Tocky Fluorescent Timer system, this study resolves signalling-linked temporal progression alongside thymocyte differentiation and shows that the learned reference structure can be transferred across species to organise human thymic single-cell data.
|
|---|---|
| Mar 13, 2026 | 📄 New Preprint Published 🚀 Our new bioRxiv preprint introduces CanonicalTockySeq, a framework that integrates Nr4a3-Tocky Fluorescent Timer signalling history with single-cell RNA-seq to resolve dynamic T-cell states in cancer immunotherapy. Using an experimentally anchored canonical manifold, this study separates temporal progression from signalling strength at single-cell resolution and shows that effective checkpoint blockade is associated with reduced persistence of antigen engagement, suppression of exhaustion-associated TCR signalling programmes, and maintenance of progenitor-like features linked to durable antitumour responses.
|
| Jul 02, 2025 | 📄 New Paper Published 🕒 Our paper in Biology Methods and Protocols introduces TockyLocus, an open-source framework for the quantitative and standardised analysis of Fluorescent Timer flow cytometry data in Nr4a3-Tocky and Foxp3-Tocky mice. By optimising analysis of Timer Angle and establishing a biologically grounded five-locus categorisation scheme, this study provides a robust and interpretable method for analysing transcriptional dynamics, enabling reproducible statistical testing and visualisation without reliance on arbitrary gating.
|
| Jul 01, 2025 | 📄 New Paper Published 🌟 Our paper in Nature Communications presents a new framework for decoding temporal transcriptional dynamics from Fluorescent Timer data using an integrative combination of molecular genetics, flow cytometry, and machine learning. Using a convolutional neural network (ConvNet)-based approach together with Foxp3-Tocky Fluorescent Timer reporter mice and CRISPR gene editing, this study reveals previously unrecognised features of Foxp3 transcriptional dynamics, including roles of CNS2 in regulating transcription frequency and age-dependent differences from neonatal to aged mice.
|
| Feb 09, 2025 | 📄 New Paper Published 🌟 Excited to share my new publication on TockyPrep, now published in BMC Bioinformatics🚀 The TockyPrep R package automates and standardizes flow cytometric analysis of Fluorescent Timer reporters, unlocking the analysis of Nr4a3 Tocky mice and other Timer reporters🐭 |
selected publications
- Canonical Analysis of Fluorescent Timer-Anchored Transcriptomes Resolves Joint Temporal and Developmental ProgressionbioRxiv, May 2026
- Temporal Mechanisms of T-Cell Fate Decisions under Immune Checkpoint Blockade Resolved by CanonicalTockySeqbioRxiv, May 2026
- Addition of chemotherapy to radiotherapy promotes progenitor-exhausted CD8+ T-cell clonal dominance in head and neck cancerbioRxiv, May 2026
- Machine learning-assisted decoding of temporal transcriptional dynamics via fluorescent timerNature Communications, May 2025Introduces TockyConvNet, a deep-learning framework that reveals enhancer-dependent and age-dependent Foxp3 transcriptional dynamics from fluorescent Timer data.
- GatingTree: Pathfinding Analysis of Group-Specific Effects in Cytometry DataCytometry Part A, May 2025Introduces GatingTree, a pathfinding framework for identifying group-specific cytometry features and deriving practical gating strategies without dimensional reduction.
- TockyLocus: quantitative analysis of flow cytometric fluorescent timer data in Nr4a3-Tocky and Foxp3-Tocky miceBiology Methods and Protocols, May 2025Establishes TockyLocus, introducing a five-locus quantitative framework that links Timer Angle geometry to interpretable transcriptional dynamics.
- TockyPrep: data preprocessing methods for flow cytometric fluorescent timer analysisBMC Bioinformatics, May 2025Establishes TockyPrep, standardizing normalization and trigonometric transformation of fluorescent Timer flow cytometry data for reproducible quantitative analysis.
- Age-dependent Zap70 expression in thymocytes regulates selection of the neonatal regulatory T cell repertoireNature Immunology, May 2025
- Xevinapant plus Chemoradiotherapy Negatively Sculpts the Tumor-Immune Microenvironment in Head and Neck CancerCancer Research Communications, Nov 2025
- Intragenic viral silencer element regulates HTLV-1 latency via RUNX complex recruitmentNature Microbiology, Nov 2025
- A multidimensional toolkit for elucidating temporal trajectories in cell development in vivoDevelopment, Nov 2024
- TockyLocus: Quantitative Analysis Methods for Flow Cytometric Fluorescent Timer DataarXiv, Nov 2024
- TockyPrep: Data Preprocessing Methods for Flow Cytometric Fluorescent Timer AnalysisarXiv, Nov 2024
- GatingTree: Pathfinding Analysis of Group-Specific Effects in Cytometry DataarXiv, Nov 2024
- Spectrum of Treg and self-reactive T cells: single cell perspectives from old friend HTLV-1Discovery Immunology, May 2024
- Control of regulatory T-cell differentiation and function by T-cell receptor signalling and Foxp3 transcription factor complexesImmunology, May 2020A cutting-edge review proposing an integrated mechanistic framework for how TCR signalling and Foxp3 complexes control regulatory T-cell differentiation and function.
- A temporally dynamic Foxp3 autoregulatory transcriptional circuit controls the effector Treg programmeThe EMBO journal, May 2018The second Tocky paper from the Ono lab uncovers the temporally dynamic regulation of Foxp3 transcription, offering new insights into T cell regulation.
- A timer for analyzing temporally dynamic changes in transcription during differentiation in vivoJournal of Cell Biology, May 2018The foundational publication introducing Tocky technology by the Ono lab, marking a breakthrough in T cell and B cell studies.
- Controversies concerning thymus‐derived regulatory T cells: fundamental issues and a new perspectiveImmunology and cell biology, May 2016A landmark opinion piece challenging the reproducibility of a foundational Treg experiment, while introducing a groundbreaking dynamic view on Foxp3-mediated T cell regulation.
- Regulatory T cells in melanoma revisited by a computational clustering of FOXP3+ T cell subpopulationsThe Journal of Immunology, May 2016The pioneering study ntroduces a data-driven methodology to explore Treg subpopulations, employing Principal Component Analysis (PCA) for the first time.
- Visualisation of the T cell differentiation programme by Canonical Correspondence Analysis of transcriptomesBMC Genomics, May 2014This study establishes CCA as a vital quantitative tool for transcriptome analysis in immunological datasets.
- Visualising the cross-level relationships between pathological and physiological processes and gene expression: analyses of haematological diseasesPLoS One, May 2013The pioneering study developing Canonical Correspondence Analysis (CCA) as a genomics method, establishing a novel approach for transcriptome analysis.

