Lab
Complementary research bases in London and Kumamoto
The Ono Lab operates across two complementary research bases in London and Kumamoto, linked by a shared interest in immune dynamics and by common experimental and computational frameworks.
The UK Lab focuses on cancer immunology and immunotherapy, with particular emphasis on how T-cell receptor signalling, Foxp3-associated programs, immune checkpoint pathways, and cytokine-responsive processes change over time in the tumour microenvironment.
The Japan Lab develops broader work on immune dynamics, with emphasis on the temporal regulation of immune responses and on the development of experimental and computational frameworks for analysing immune cell state transitions.
UK Lab (London)
The UK Lab is centred on cancer immunology and immunotherapy. Its work examines how immune cell states emerge, persist, and change over time during anti-tumour responses and immune checkpoint blockade.
Cancer Immunology and Immunotherapy
Our research focuses on cancer immunology and immunotherapy, with particular emphasis on understanding immune responses as dynamic processes evolving over time rather than as static cell categories.
In particular, we study how T-cell receptor signalling, Foxp3-associated programs, immune checkpoint pathways, and cytokine-responsive processes are regulated, integrated, and reshaped over time in the tumour microenvironment. This is addressed through a combination of experimental immunology and computational analysis.
Building on the Tocky system based on Fluorescent Timer proteins, we aim to capture not only the current state of immune cells, but also their recent stimulation history and the trajectory of their responses during anti-tumour immunity and immunotherapy.
Current research themes include:
- temporal mechanisms of T-cell fate in cancer
- immune checkpoint biology
- immunotherapy response trajectories
- Foxp3-associated regulatory programs in tumour immunity
- time-resolved single-cell analysis of tumour-infiltrating immune cells

Japan Lab (Kumamoto)
The Japan Lab, 免疫・感染ダイナミックス分野 at Kumamoto University, provides a broader base for studying immune dynamics in vivo.
Its work focuses on the dynamic regulation of immune cell states, signalling history, and gene regulatory programs through the integration of experimental immunology and computational analysis. This includes the development of Tocky-based approaches and related analytical frameworks for resolving immune responses over time.

Research Overview
Anti-tumour immunity is shaped by continuously changing cellular states in the tumour microenvironment.
Our work focuses on the temporal regulation of immune responses in cancer, aiming to understand state transitions, differentiation processes, adaptation to persistent stimulation, immune checkpoint-mediated regulation, and gene expression control over time.
Major areas of interest include cancer immunology, immunotherapy, immune checkpoint biology, Foxp3-associated regulatory programs, and dynamic immune responses in tumour settings.
Research Themes
Temporal Dynamics of T-cell Receptor Signalling in Cancer
We analyse how T cells respond to tumour-associated antigen stimulation in terms of both intensity and duration. Rather than treating activation as a binary state, we aim to understand it as a continuous temporal process within the tumour microenvironment.
Immune Checkpoint Pathways and Immunotherapy Response
We investigate how immune checkpoint pathways are induced and regulated over time, and how they influence immune cell states during checkpoint blockade and related immunotherapies.
Dynamics of Foxp3 and Immune Regulatory Molecules in Tumours
We study how Foxp3 expression, immune checkpoint molecules, and cytokine-related pathways are induced, maintained, and modulated under TCR signalling and other signalling inputs in tumour-associated immune cells. Our goal is to understand immune regulation as a dynamic molecular network rather than a fixed classification of cell types.
Immune Cell State Transitions in the Tumour Microenvironment
We aim to characterise how immune cells change over time in tumours, and to understand both productive anti-tumour responses and dysfunctional or suppressed states.
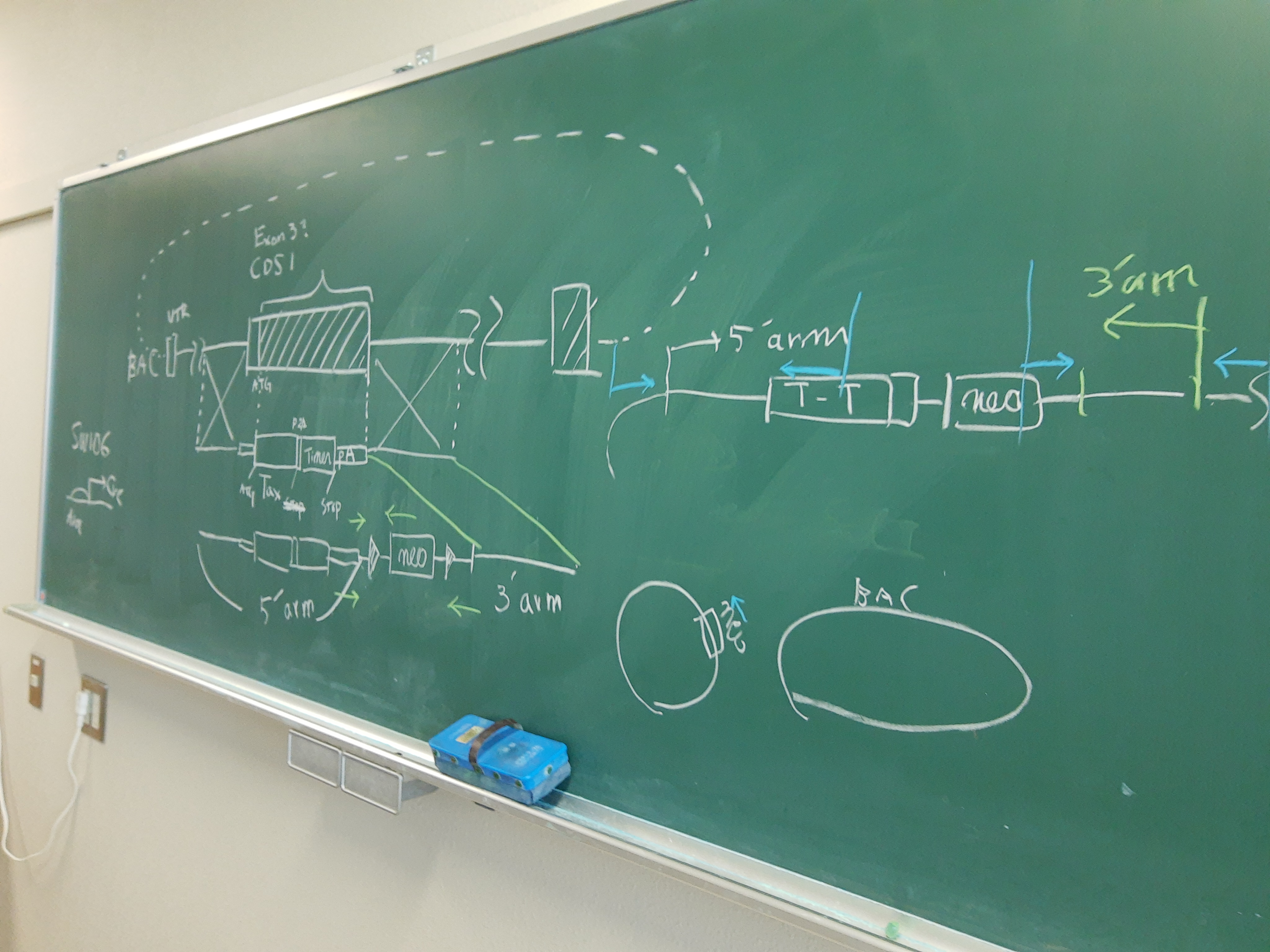
Technical Platforms
Tocky
We use Fluorescent Timer proteins to visualise temporal information in intracellular events and immune signalling dynamics.
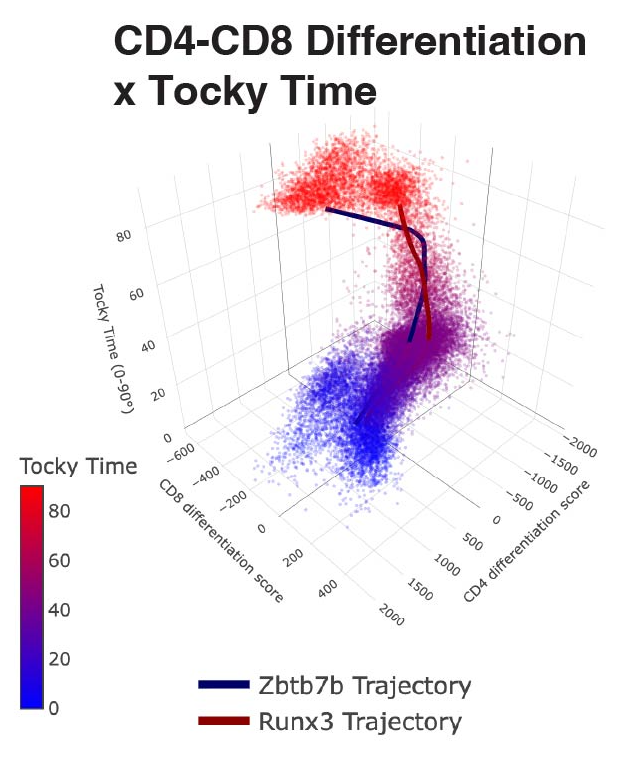
Figure from our bioRxiv preprint: Canonical Analysis of Fluorescent Timer-Anchored Transcriptomes Resolves Joint Temporal and Developmental Progression.
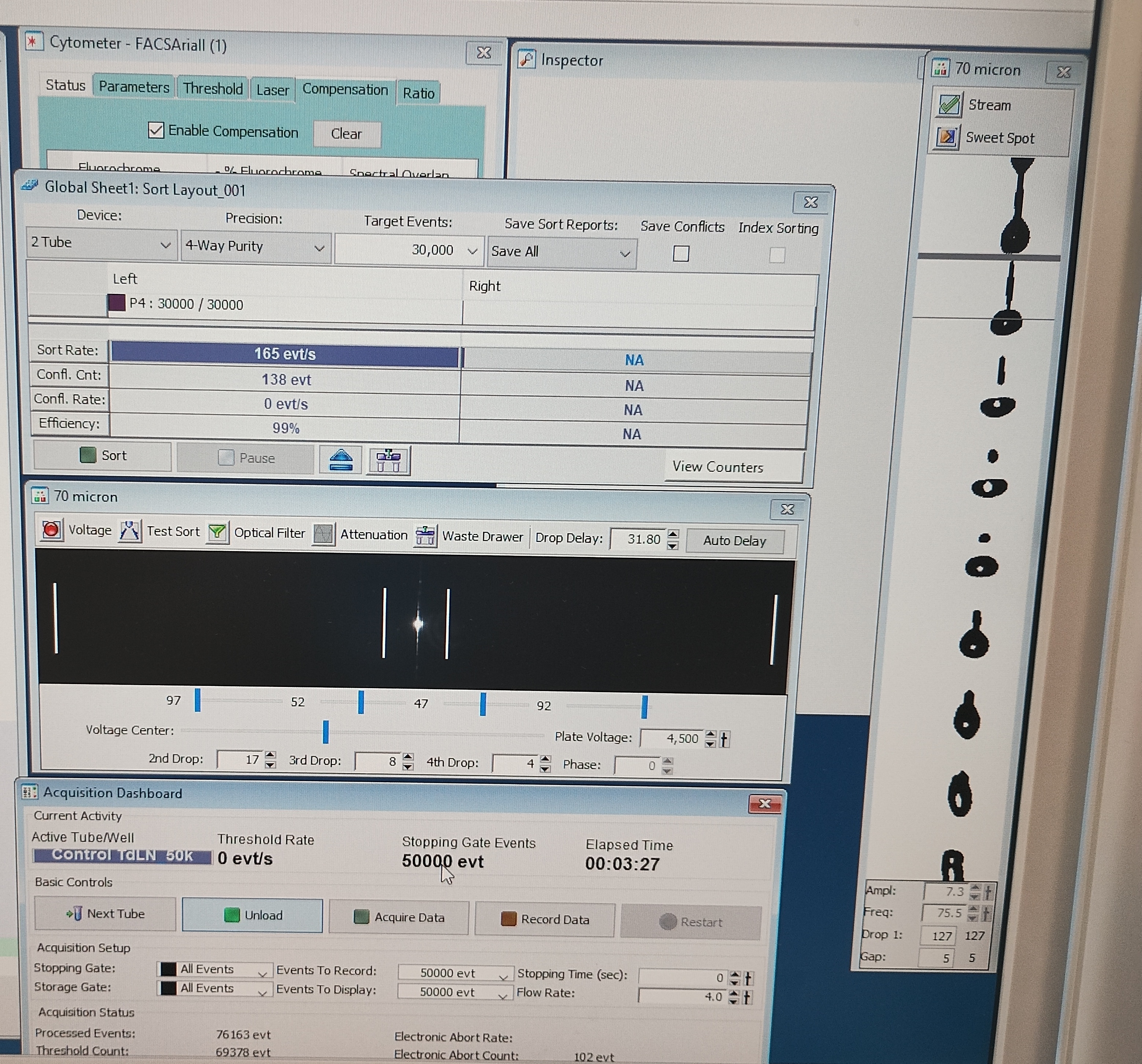
Flow Cytometry
We use high-dimensional flow cytometry to analyse immune cell states at high resolution.
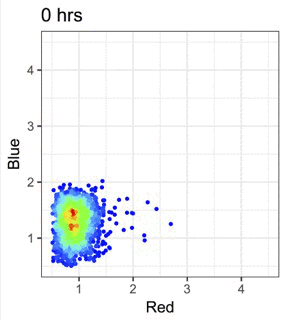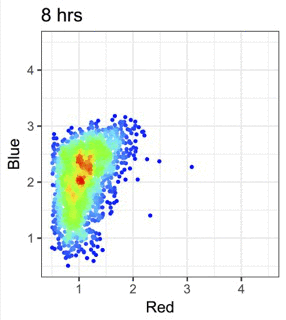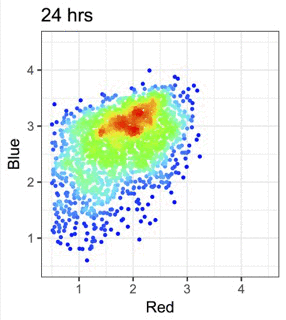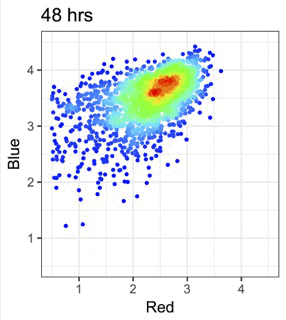
Single-Cell Analysis
We analyse immune cell heterogeneity and state transitions using single-cell transcriptomics and flow cytometry.
Integration of Machine Learning and Immune Data Analysis
We develop analytical methods for Tocky and flow cytometry data using machine learning and multidimensional analysis. These methods are shared broadly to support the development of a robust analytical ecosystem and research infrastructure for dynamic immune profiling.
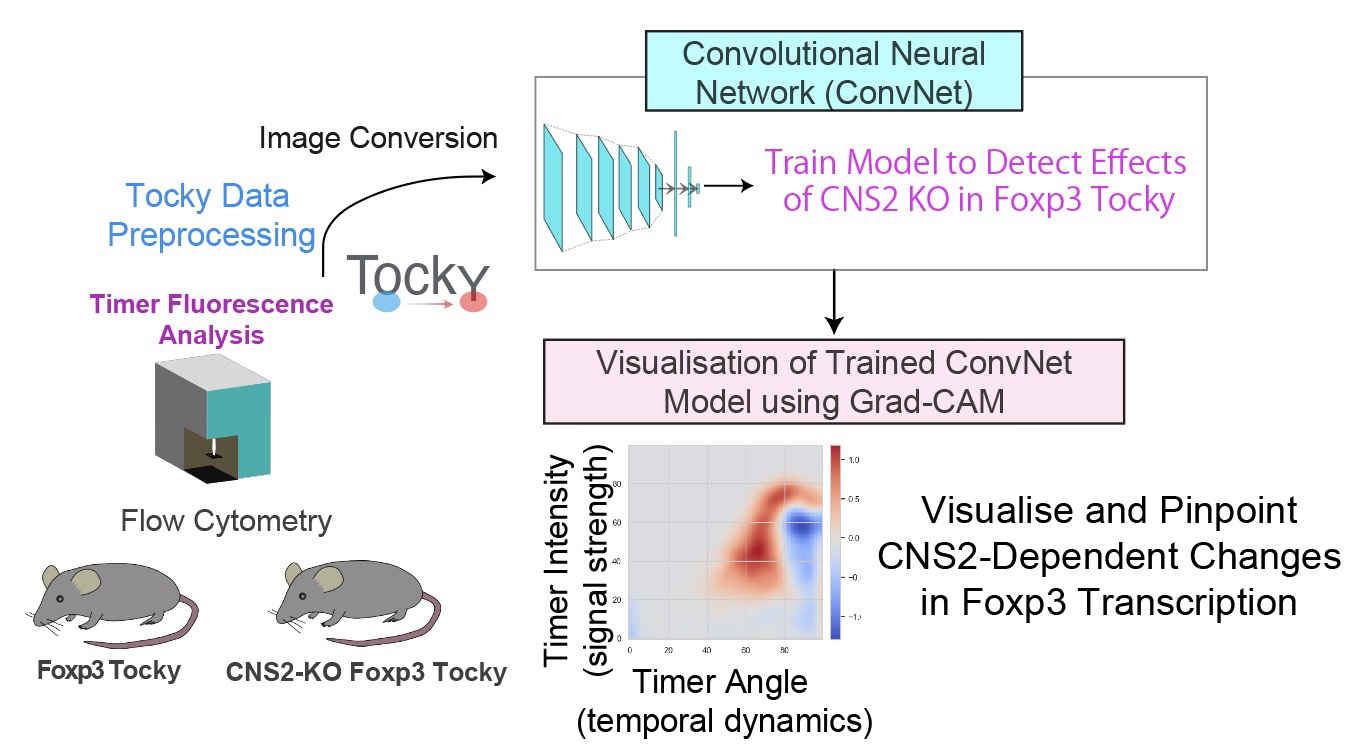
Figure modified from Irie et al., Nature Communications (2025): Machine learning-assisted decoding of temporal transcriptional dynamics via fluorescent timer.
Collaboration and Opportunities
We welcome collaborative research in cancer immunology, immunotherapy, flow cytometry, single-cell analysis, and computational approaches to dynamic immune profiling.
We also welcome students and early-career researchers interested in combining experimental biology with quantitative and data-driven analysis.
Contact
news
| Mar 18, 2026 | 📄 New Preprint Published 🧭 Our new bioRxiv preprint introduces mCanonicalTockySeq, a framework that brings an experimental time anchor into single-cell transcriptomics to analyse temporal and developmental progression together. Using the Nr4a3-Tocky Fluorescent Timer system, this study resolves signalling-linked temporal progression alongside thymocyte differentiation and shows that the learned reference structure can be transferred across species to organise human thymic single-cell data.
|
|---|---|
| Mar 13, 2026 | 📄 New Preprint Published 🚀 Our new bioRxiv preprint introduces CanonicalTockySeq, a framework that integrates Nr4a3-Tocky Fluorescent Timer signalling history with single-cell RNA-seq to resolve dynamic T-cell states in cancer immunotherapy. Using an experimentally anchored canonical manifold, this study separates temporal progression from signalling strength at single-cell resolution and shows that effective checkpoint blockade is associated with reduced persistence of antigen engagement, suppression of exhaustion-associated TCR signalling programmes, and maintenance of progenitor-like features linked to durable antitumour responses.
|
| Jul 02, 2025 | 📄 New Paper Published 🕒 Our paper in Biology Methods and Protocols introduces TockyLocus, an open-source framework for the quantitative and standardised analysis of Fluorescent Timer flow cytometry data in Nr4a3-Tocky and Foxp3-Tocky mice. By optimising analysis of Timer Angle and establishing a biologically grounded five-locus categorisation scheme, this study provides a robust and interpretable method for analysing transcriptional dynamics, enabling reproducible statistical testing and visualisation without reliance on arbitrary gating.
|
| Jul 01, 2025 | 📄 New Paper Published 🌟 Our paper in Nature Communications presents a new framework for decoding temporal transcriptional dynamics from Fluorescent Timer data using an integrative combination of molecular genetics, flow cytometry, and machine learning. Using a convolutional neural network (ConvNet)-based approach together with Foxp3-Tocky Fluorescent Timer reporter mice and CRISPR gene editing, this study reveals previously unrecognised features of Foxp3 transcriptional dynamics, including roles of CNS2 in regulating transcription frequency and age-dependent differences from neonatal to aged mice.
|
| Feb 09, 2025 | 📄 New Paper Published 🌟 Excited to share my new publication on TockyPrep, now published in BMC Bioinformatics🚀 The TockyPrep R package automates and standardizes flow cytometric analysis of Fluorescent Timer reporters, unlocking the analysis of Nr4a3 Tocky mice and other Timer reporters🐭 |
selected publications
- Canonical Analysis of Fluorescent Timer-Anchored Transcriptomes Resolves Joint Temporal and Developmental ProgressionbioRxiv, 2026
- Temporal Mechanisms of T-Cell Fate Decisions under Immune Checkpoint Blockade Resolved by CanonicalTockySeqbioRxiv, 2026
- Addition of chemotherapy to radiotherapy promotes progenitor-exhausted CD8+ T-cell clonal dominance in head and neck cancerbioRxiv, 2026
- Machine learning-assisted decoding of temporal transcriptional dynamics via fluorescent timerNature Communications, 2025Introduces TockyConvNet, a deep-learning framework that reveals enhancer-dependent and age-dependent Foxp3 transcriptional dynamics from fluorescent Timer data.
- GatingTree: Pathfinding Analysis of Group-Specific Effects in Cytometry DataCytometry Part A, 2025Introduces GatingTree, a pathfinding framework for identifying group-specific cytometry features and deriving practical gating strategies without dimensional reduction.
- TockyLocus: quantitative analysis of flow cytometric fluorescent timer data in Nr4a3-Tocky and Foxp3-Tocky miceBiology Methods and Protocols, 2025Establishes TockyLocus, introducing a five-locus quantitative framework that links Timer Angle geometry to interpretable transcriptional dynamics.
- TockyPrep: data preprocessing methods for flow cytometric fluorescent timer analysisBMC Bioinformatics, 2025Establishes TockyPrep, standardizing normalization and trigonometric transformation of fluorescent Timer flow cytometry data for reproducible quantitative analysis.
- Age-dependent Zap70 expression in thymocytes regulates selection of the neonatal regulatory T cell repertoireNature Immunology, 2025
- Xevinapant plus Chemoradiotherapy Negatively Sculpts the Tumor-Immune Microenvironment in Head and Neck CancerCancer Research Communications, Nov 2025
- Intragenic viral silencer element regulates HTLV-1 latency via RUNX complex recruitmentNature Microbiology, Nov 2025
- A multidimensional toolkit for elucidating temporal trajectories in cell development in vivoDevelopment, Nov 2024
- TockyLocus: Quantitative Analysis Methods for Flow Cytometric Fluorescent Timer DataarXiv, Nov 2024
- TockyPrep: Data Preprocessing Methods for Flow Cytometric Fluorescent Timer AnalysisarXiv, Nov 2024
- GatingTree: Pathfinding Analysis of Group-Specific Effects in Cytometry DataarXiv, Nov 2024
- Spectrum of Treg and self-reactive T cells: single cell perspectives from old friend HTLV-1Discovery Immunology, May 2024
- Control of regulatory T-cell differentiation and function by T-cell receptor signalling and Foxp3 transcription factor complexesImmunology, May 2020A cutting-edge review proposing an integrated mechanistic framework for how TCR signalling and Foxp3 complexes control regulatory T-cell differentiation and function.
- A temporally dynamic Foxp3 autoregulatory transcriptional circuit controls the effector Treg programmeThe EMBO journal, May 2018The second Tocky paper from the Ono lab uncovers the temporally dynamic regulation of Foxp3 transcription, offering new insights into T cell regulation.
- A timer for analyzing temporally dynamic changes in transcription during differentiation in vivoJournal of Cell Biology, May 2018The foundational publication introducing Tocky technology by the Ono lab, marking a breakthrough in T cell and B cell studies.
- Controversies concerning thymus‐derived regulatory T cells: fundamental issues and a new perspectiveImmunology and cell biology, May 2016A landmark opinion piece challenging the reproducibility of a foundational Treg experiment, while introducing a groundbreaking dynamic view on Foxp3-mediated T cell regulation.
- Regulatory T cells in melanoma revisited by a computational clustering of FOXP3+ T cell subpopulationsThe Journal of Immunology, May 2016The pioneering study ntroduces a data-driven methodology to explore Treg subpopulations, employing Principal Component Analysis (PCA) for the first time.
- Visualisation of the T cell differentiation programme by Canonical Correspondence Analysis of transcriptomesBMC Genomics, May 2014This study establishes CCA as a vital quantitative tool for transcriptome analysis in immunological datasets.
- Visualising the cross-level relationships between pathological and physiological processes and gene expression: analyses of haematological diseasesPLoS One, May 2013The pioneering study developing Canonical Correspondence Analysis (CCA) as a genomics method, establishing a novel approach for transcriptome analysis.

